Senescence-Methylation-Signature - Determination of cellular aging in cell cultures
Ref.-Nr. 2910
Keywords: Cell culture, long-term culture, quality control, cells, cellular aging, senescence, epigenetics, DNA-methylation
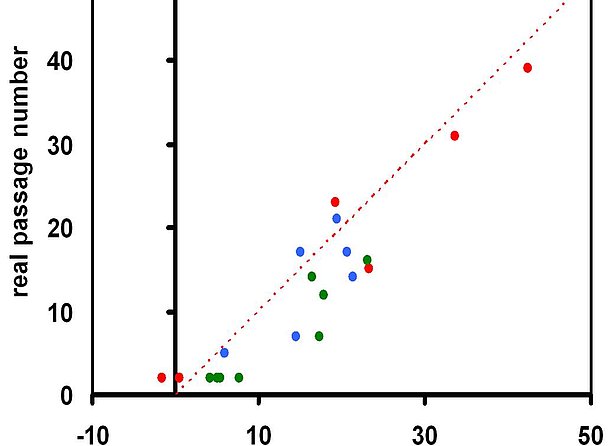
Long-term culture associated changes need to be considered for quality control of cell preparations. The invention discloses a simple method to track cellular aging based on continuous DNA-methylation changes at six specific CpG sites. This epigenetic signature can be used as a biomarker for various cell types to predict – continuously over the time – the state of cellular aging with regard to the number of passages or days of in vitro culture.
Vorteile
- Epigenetic-Aging-Signature
- Suitable for: Forensic Analysis, Geriatric assessment, Lifestyle Management
- Cost effective and simple
- Precise: Less than five years deviation
Kommerzielle Anwendung
Cross-contaminated and misidentified cell lines are a major problem in cell biology. In analogy, cross-contamination or mixing up of donor samples are also problematic for primary cells. Quality control is even more important for cellular therapy – the use of early passages is usually recommended due to less functional changes and less accumulation of mutations. Scientists of the RWTH Aachen developed a simple epigenetic test to track cellular aging. The method does not require a reference-sample of the corresponding early passage and provides a simple quality control to determine the state of cellular aging. As induced pluripotent stem cells (iPS) do not reveal cellular aging and are predicted passage zero, the identification of high quality induced iPS might also be possible. On behalf of RWTH Aachen, PROvendis offers access to rights for commercial use as well as the opportunity for further co-development.
Aktueller Stand
In case of interest we are pleased to inform you about the current patent status.
Relevante Veröffentlichungen
Koch, C.M., et al. (2012) Monitoring of cellular senescence by DNA-methylation at specific CpG sites. Aging Cell 11: 366-9.
Schellenberg, A., et al. (2014) Proof of Principle: Quality Control of Therapeutic Cell Preparations using Senescence-Associated DNA-Methylation Changes. BMC Research Notes 7: 254.
—
Eine Erfindung der RWTH Aachen.